The past couple of weeks have involved some steep learning curves. For one project, I’m running analyses in RevBayes, a very powerful Bayesian platform for doing all sorts of things related to phylogenetics. The two analyses I’m running are a divergence time estimate, and an ancestral biogeography, in a script that simultaneously infers the phylogeny of the tree. My data set is 147 cacti, and an alignment of more than 73,000 nucleotides. I love RevBayes but I have to say it’s sometimes… love hate. For one thing, working with the code is very challenging for me. I am not a natural when it comes to code. For another thing, you can’t realistically run these analyses on your home computer, so, connected with getting these results is the labyrinthine tasks involved in getting a job up and running on the High Performance Computing cluster at U of A. In fact, even there, the maximum wall time is 10 days, which is about half what I need, so I got some help from a hero in IT, running the analyses on a remote computer with unlimited time for the job to run.
But both of these tasks has required learning my way around the terminal in ways I have just never learned before.
How hard could it be?
well pretty damned hard lol. In addition to learning how to submit a batch job to the cluster, I have learned how to start a job in tmux and have it keep running whether I’m logged in or not, in a docker container. Literally a week ago I had no idea what any of that meant.
Also in addition, I have wanted to get faster results for time-calibrated phylogenies for the group I’m working on, so I have had to learn how to run analyses in treePL, which is cool, but getting even an easy start was not….easy.
I have triumphed over all of this, however. Now, the next giant leap is learning how to develop the model inputs for an Integral Projection Model (IPM), thanks to two experts from Mexico with whom I’m working on another project. I experimented with IPMs during my PhD, seven years ago or more, and the landscape has totally changed. The package I was working in doesn’t even really exist anymore.
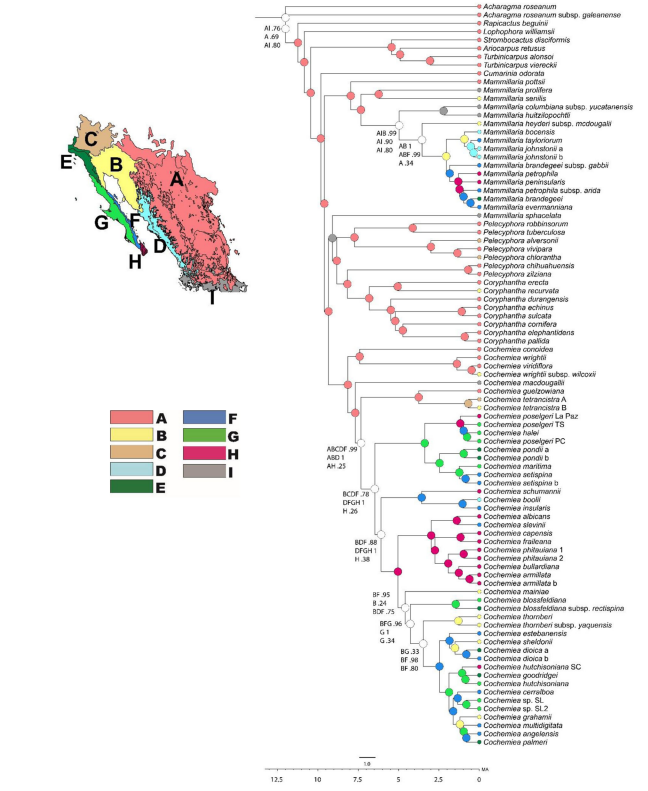
But here’s a cool figure from a paper we published a few years ago which relied very heavily on RevBayes. The figure presents the results of our ancestral biogeography analysis for some cool cacti. You can trace the wandering clades by comparing to the color-coded map and key.
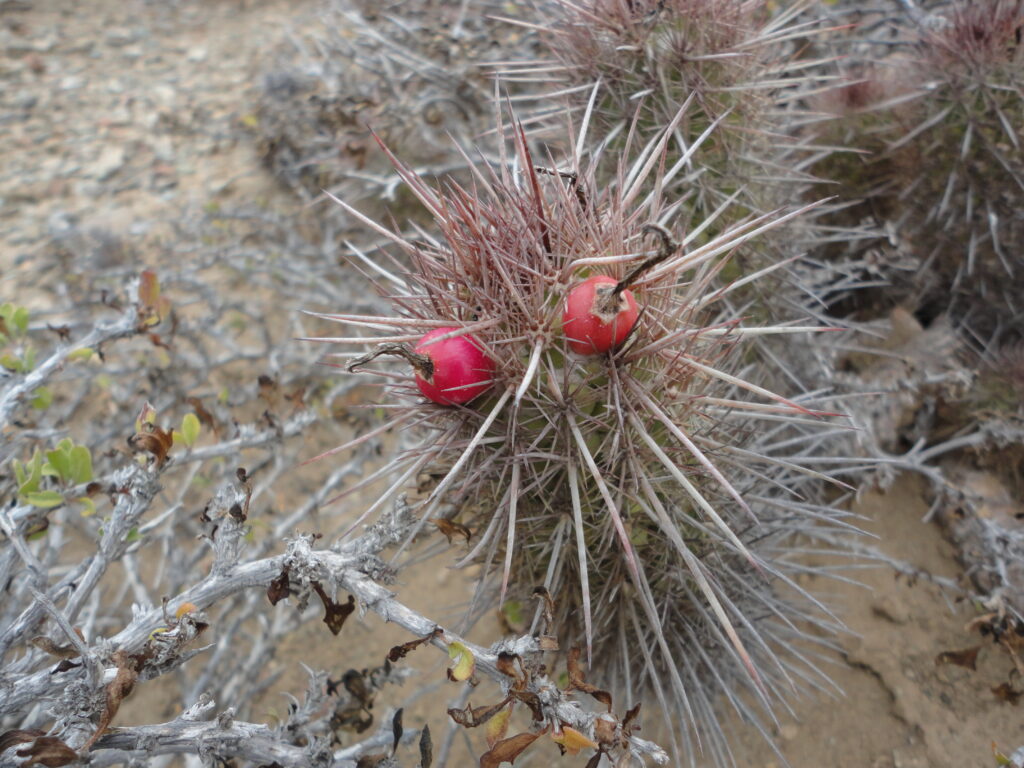

Here’s one of those cool cacti, Cochemiea halei, in habitat on a remote desert island in the Pacific Ocean.
Anyway, as fried as my brain feels after two weeks of digital madness, I wouldn’t trade it for anything. There’s nothing quite like finally running an analysis that gives you solid and reportable results. No matter how much you curse at the sinister RevBayes along the way.
